plotGrouper 
by John D. Gagnon
University of California, San
Francisco
Table of Contents
Overview
Installation
Usage
Session info
License
Overview
A shiny app-based GUI wrapper for ggplot2 with built-in statistical
analysis. Import data from file and use dropdown menus and checkboxes to
specify the plotting variables, graph type, and look of your plots. Once
created, plots can be saved independently or stored in a report that can
be saved as a pdf. If new data are added to the file, the report can be
refreshed to include new data. Statistical tests can be selected and
added to the graphs.
Analysis of flow cytometry data is especially integrated with
plotGrouper. Count data can be transformed to return the absolute number
of cells in a sample (this feature requires inclusion of the number of
beads per sample and information about any dilution performed).
Examples of some of the types of plots you can create:
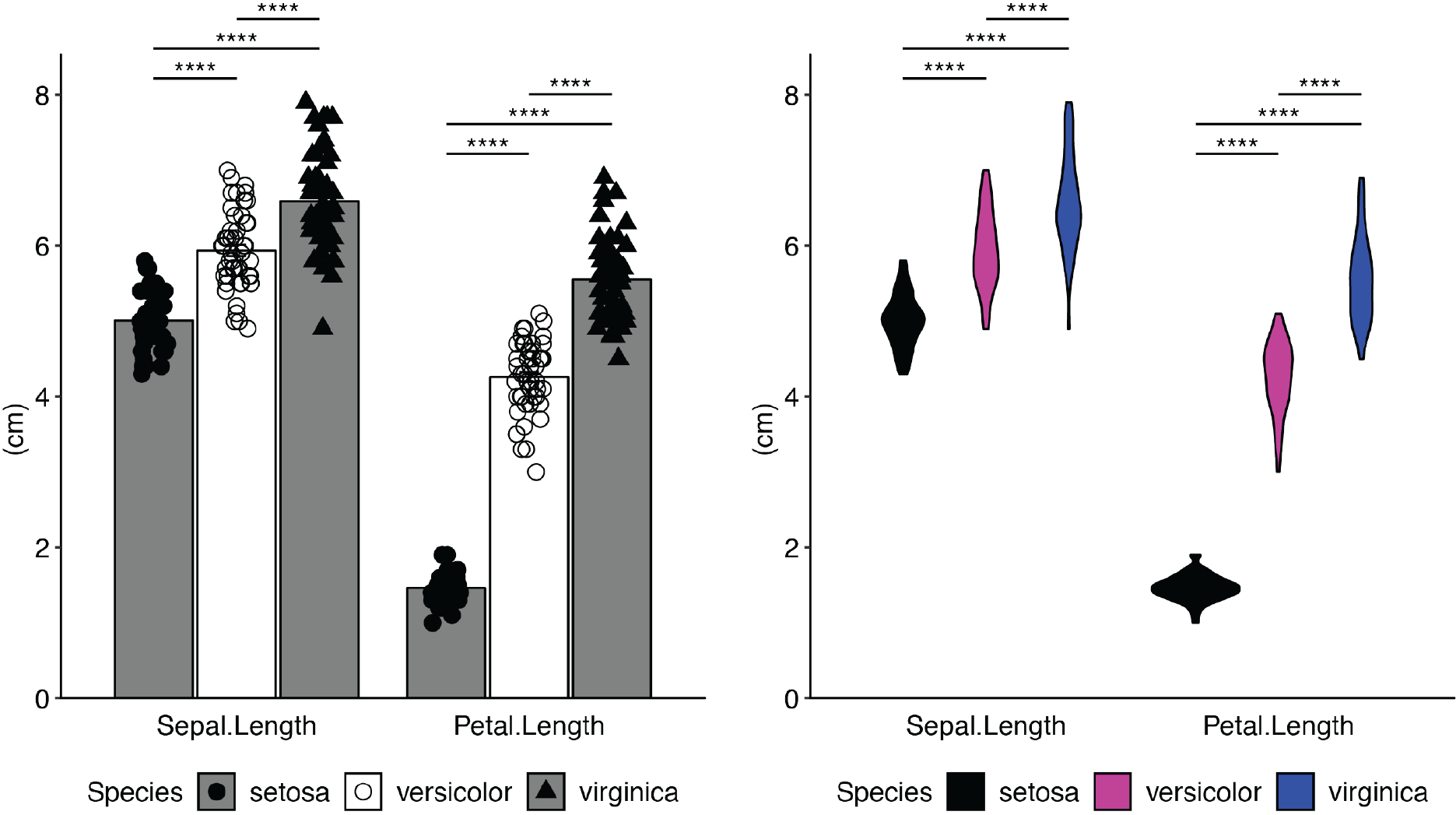
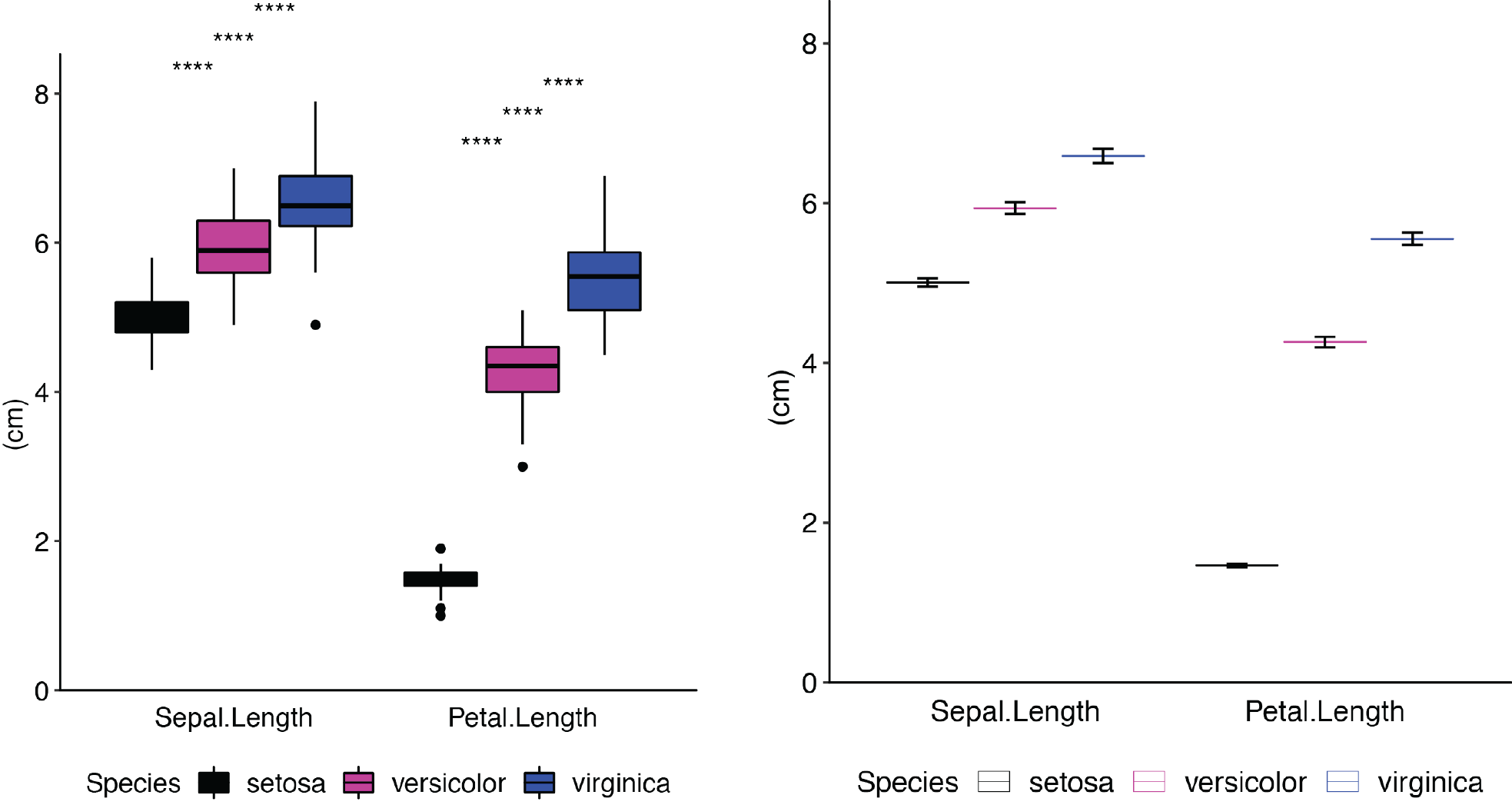
Installation
-
If you do not already have R installed, or your version is out of date, download and install the latest version.
- Optionally, install the latest version of RStudio Desktop.
-
Download the package from Bioconductor.
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("plotGrouper")
- Or install the development version of the package from Bioconductor:
'Bioconductor'
BiocManager::install("plotGrouper", version = "devel")
- Or GitHub:
BiocManager::install("jdgagnon/plotGrouper")
Usage
Load the package into the R session.
library(plotGrouper)
To initialize the shiny app, paste the following code in your R console and run it.
plotGrouper()
Once the web app opens, you can access the iris dataset by clicking
the iris button to learn how to use the app. After the iris data
loads, the selection windows will be automatically populated and a graph
should be displayed.
The Raw Data tab displays the structure of the data loaded. Your file
should be organized in the following way:
| Unique identifier | Comparisons | Variables |
|---|---|---|
| Sample | Species | Sepal.Length |
| setosa_1 | setosa | 5.1 |
| setosa_2 | setosa | 4.9 |
| versicolor_1 | versicolor | 7 |
| versicolor_2 | versicolor | 6.4 |
| virginica_1 | virginica | 6.3 |
| virginica_2 | virginica | 5.8 |
| etc… | etc… | etc… |
These columns can be titled anything you want but values in the columns are important.
-
The
Unique identifiercolumn should contain only unique values that identify each individual sample (e.g.,SamplewithinirisRaw Data). -
The
Comparisonscolumn should contain replicated values that identify each individual as belonging to a group (e.g.,SpecieswithinirisRaw Data). -
The
Variablescolumn(s) should created for each variable you wish to plot. The values in these columns must be numeric (e.g.,Sepal.Length,Sepal.Width,Petal.Length,Petal.WidthwithinirisRaw Data)
After importing a data file, a Sheet column will be created and
populated with the sheet name(s) from the file if it came from an excel
spreadsheet or the file name if it came from a csv or tsv file.
-
The
Variables to plotselection window is used to choose which variable(s) to plot (e.g.,Sepal.Widthfrom theirisdata). If multiple are selected, they will be grouped according to theIndependent variableselected. -
The
Comparisonsselection window is used to choose which column contains the information that identifies which condition each sample belongs to (e.g., theSpeciescolumn within theirisdata). -
The
Independent variableselection window is used to select how the plots should be grouped. Ifvariableis selected (the default), the plots will be grouped by the values inVariables to plot. -
Use the
Shapesselector to change the shape of the points for each comparison variable. -
Use the
Colorsselector to change the point colors for each comparison variable. -
Use the
Fillsselector to change the fill color for the other geoms being plotted for each comparison variable.
To prevent the Shapes, Colors, or Fills from reverting to their
defaults, click the Lock checkboxes.
Individual plots can be saved by clicking Save on the Plot tab or
multiple plots may be arranged on a single page by clicking Add plot to
report. Clicking this button will send the current plot to the Report
tab and assign it a number in the Report plot # dropdown menu. To
revisit a plot stored in the Report tab, select the plot you wish to
restore and click Load plot from report. Changes can be made to this
plot and then updated in the Report by clicking Update plot in
report.
-
The statistics calculated for the current plot being displayed in the
Plottab are stored in theStatisticstab. These can be saved by clicking theDownloadbutton on theStatisticstab. -
The
Plot Datatab contains the reorganized subset of data being plotted. -
The
Raw Datatab displays the dataframe that was created upon import of the file along with the automatically createdSheetcolumn.
Session info
Here is the output of sessionInfo() on the system on which this
package was developed:
sessionInfo()
#> R version 3.5.1 (2018-07-02)
#> Platform: x86_64-apple-darwin17.6.0 (64-bit)
#> Running under: macOS High Sierra 10.13.6
#>
#> Matrix products: default
#> BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
#> LAPACK: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libLAPACK.dylib
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> loaded via a namespace (and not attached):
#> [1] compiler_3.5.1 backports_1.1.2 magrittr_1.5 rprojroot_1.3-2
#> [5] tools_3.5.1 htmltools_0.3.6 yaml_2.2.0 Rcpp_0.12.19
#> [9] stringi_1.2.4 rmarkdown_1.10 highr_0.7 knitr_1.20
#> [13] stringr_1.3.1 digest_0.6.18 evaluate_0.12